Molecular Fluid Dynamics
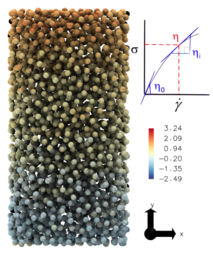
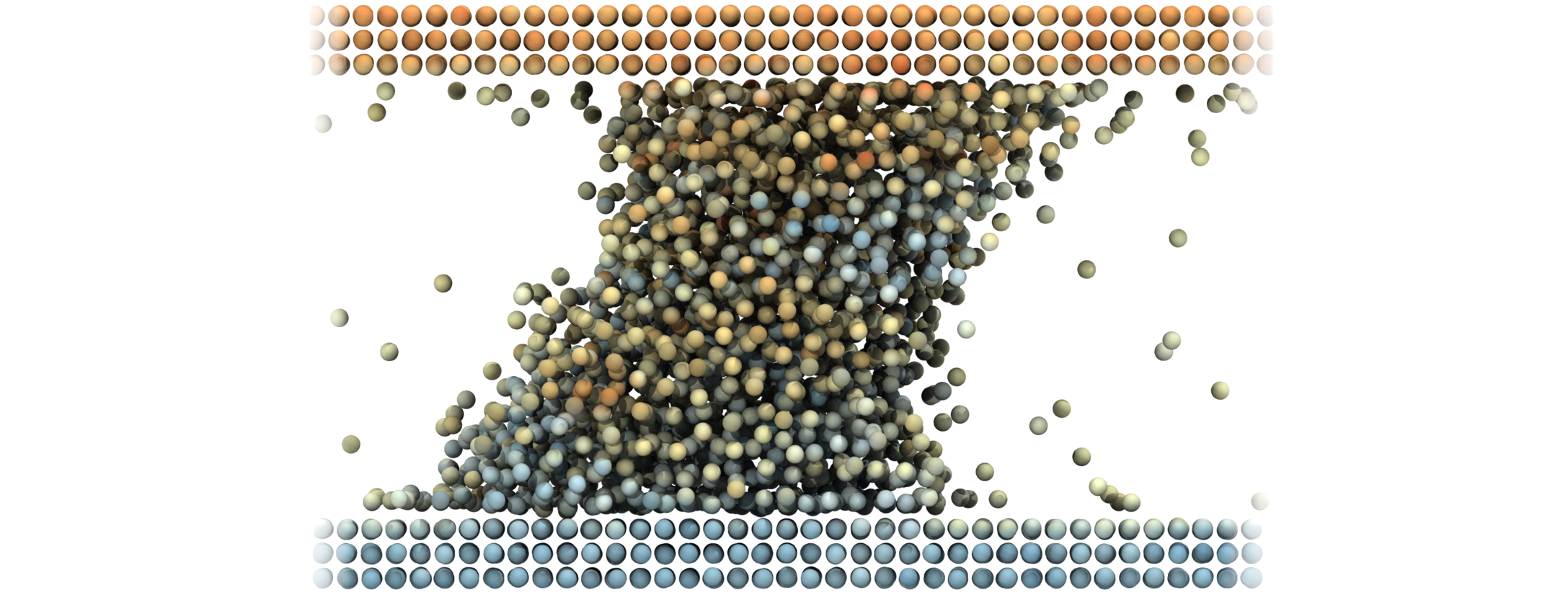
I use Molecular dynamics to simulate fluid flow, including the first ever simulation of turbulence at the molecular scale, the fluid-vapour interface developing an exact control volume framework , modelling heat flux beyond Fourier's law and studying the moving contact line. Molecular fluid dynamics is often known as Non-equilibrium molecular dynamics (NEMD) and I have worked extensively in the NEMD community, leading the UK fluids network special interest group (SIG), and together with collaborators worked on the definition of stress , Tribological properties in a molecular system, extraction of order from chaotic molecular systems and developed a new analysis studying a quantity called the viscuit .
Coupled Simulation
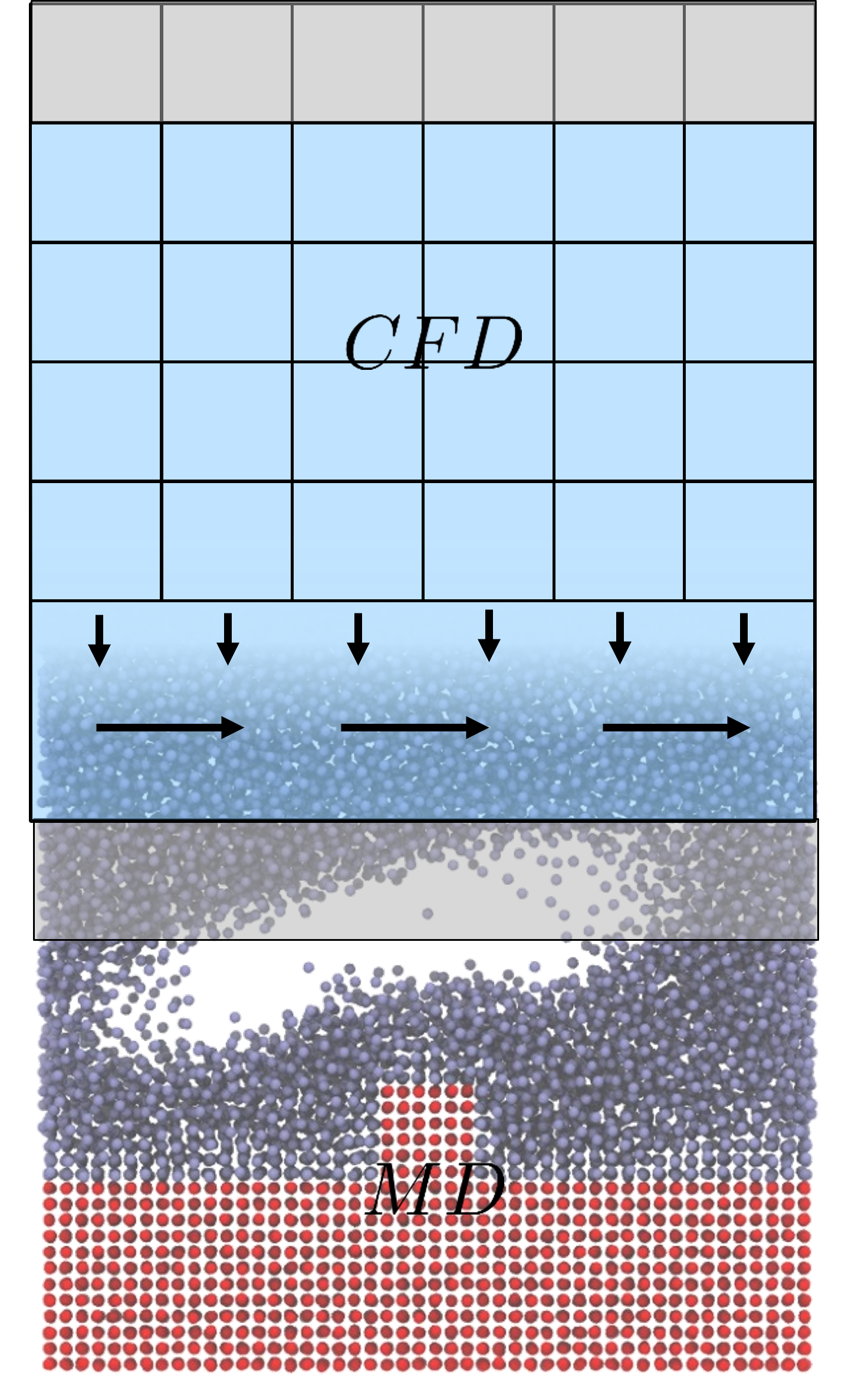
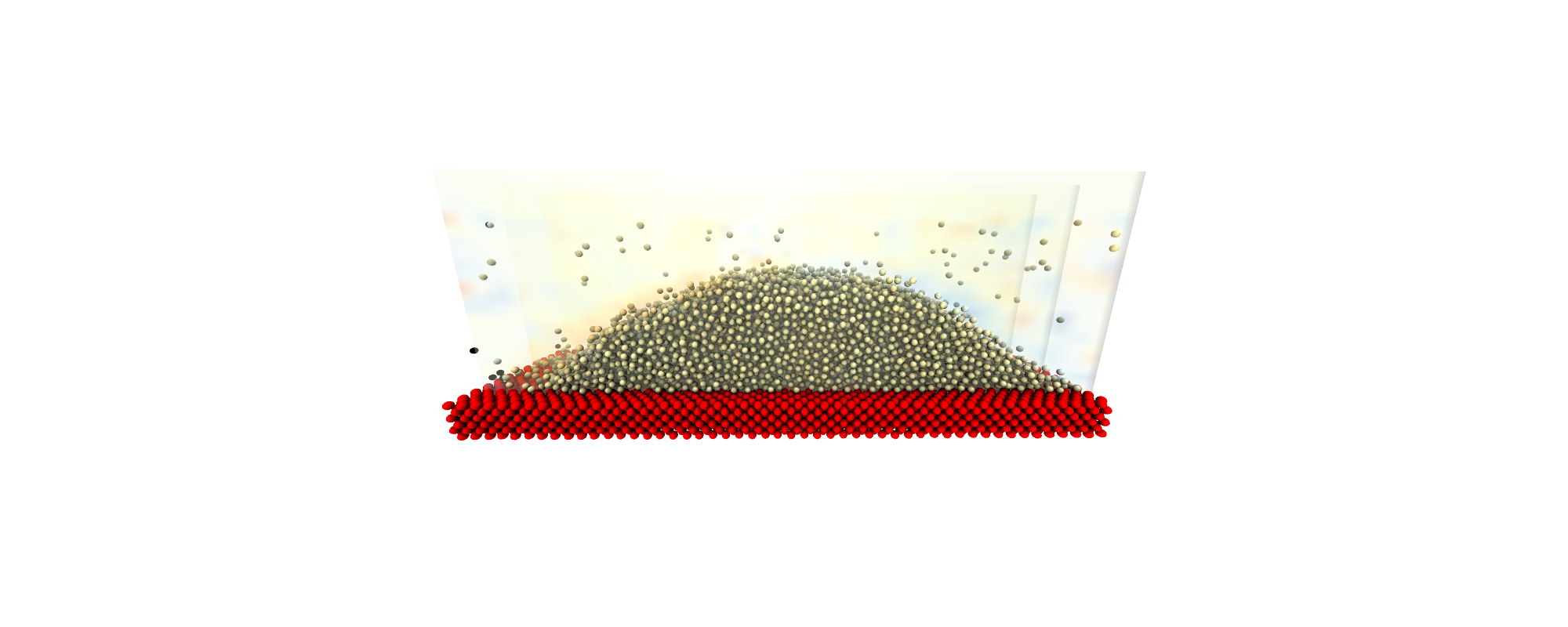
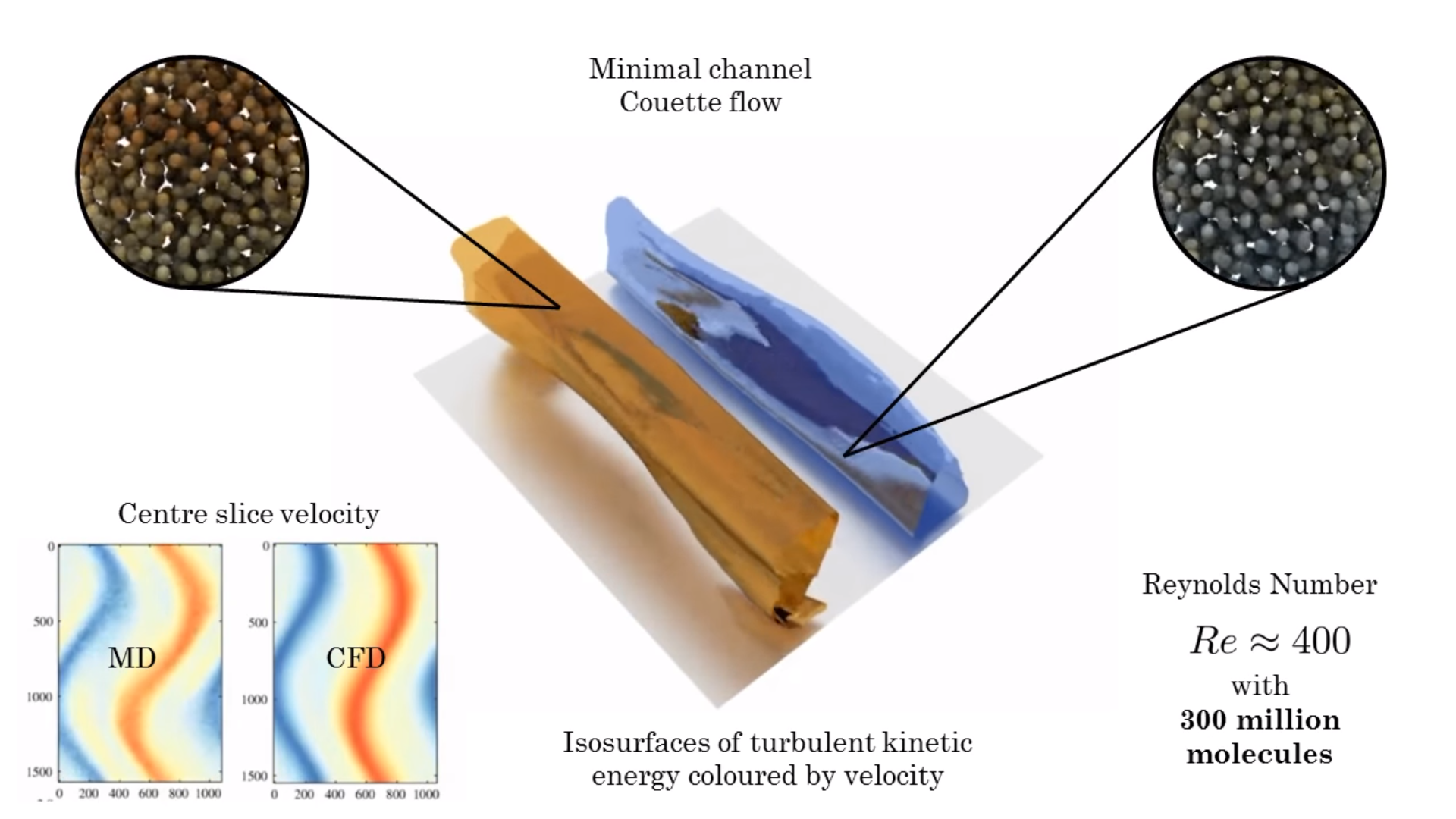
I work to develop a mathematical framework for multi-scale coupled simulation of molecular dynamics and computational fluid dynamics, including an extension of the control volume formulation to discrete system and a constrained dynamics methodology to allow exact conservation of momentum in a molecular system. Combining these development, I aim to give coupled simulation a rigourous theoretical underpinning with open-source algorithms available in my molecular dynamics code, FlowMol and coupling through CPL library website ( www.cpl-library.org ). Example application include coupled turbulence and coupled bubble nucleation .
Software Development
My research is based on my own molecular dynamics code, FlowMol , which is highly parallelised for hundreds of cores of an HPC platforms and open source with a (currently experimental) GUI. I am passionate about best practice in software, leading the development of coupling framework CPL library organising training for the Materials and Molecular Modelling Hub, a 14 university consortium linked to the supercomputer Young and teaching programming at Brunel University. I'm a fellow of the software sustainability institute SSI , created a new Python course at Imperial, work for open research through software as explained in this open research award interview video and answer questions on stackoverflow .